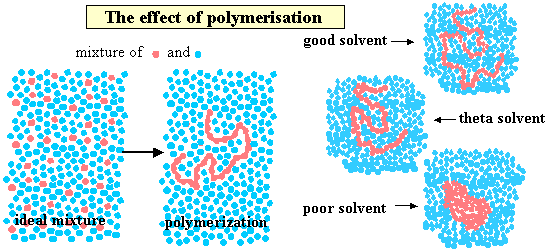
A polymer is generated by joining many monomers together. The figure below shows a monomer species (left) and the corresponding polymer (right) in solution, where the monomer solution is ideal. The segments of the polymer are restricted in comparison with the free monomer. This has consequences for both the dynamics and the thermodynamics of the solution. One consequence is that any unfavourable interaction of the monomer unit with the solvent is magnified by the large number of such interactions for each polymer molecule. Since dissimilarity generally leads to a positive enthalpy of mixing this means that the polymer only dissolves in a solvent with which its monomer is closely similar. Thus, it is common that solvents that dissolve the monomer do not dissolve the polymer. The right hand side of the figure shows what happens to the polymer dimensions as the "quality" of the solvent is varied from poor (polymer solubility reduced), through the θ condition (where there is no nett effect of solvent on the polymer dimensions) to good.

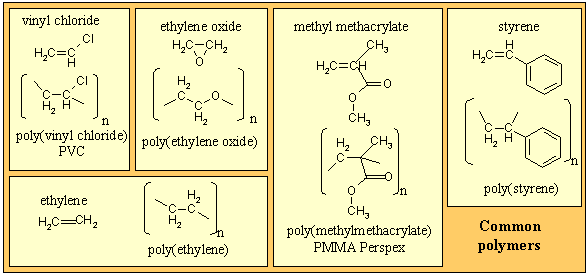
There are many types of polymer, the simplest being a linear connection of one chemical species to form a homopolymer, of which several examples are given in the panel below. Copolymers involve more than one chemical species and they can be combined in a variety of ways, e.g. alternating, randomly or in blocks. There are also non-linear combinations, e.g. star, comb, dendritic, etc. The selective use of these structural possibilities leads to a wide range of technological applications of polymeric materials.
The simplest model of a polymer structure is the three dimensional random walk (random flight; the next section describes this in detail). The applet below ( click here for notes about the use of java applets and click here for other physical chemistry applets; for downloading the relevant filename for the polymer applet below is randWalk.jar and the code to operate it is given at the end of this document) calculates a new one thousand step random walk each time the button is pressed. The end to end distance is given together with its cumulative average.
The polymer consists, at start-up, of 1000 segments of length unity and these are free to rotate through any angle with respect to adjacent segments (random flight model, three dimensional random walk model). The polymer is drawn in perspective to capture the 3-D structure. The two ends of the polymer are marked with black rings. The brighter coloured, larger segments are in the foreground and the paler, smaller segments are furthest away. On pressing New random walk a new configuration is generated. The right hand side shows the square root of the end-to-end distance of the configuration shown and the accumulated root mean square end-to-end distance. It also plots a histogram of the accumulated RMS end-to-end distances. There is one adjustable parameter, which is discussed further below.
You should try the following.
(i) Repeat the calculation enought times to generate a reasonably stable histogram of RMS end-to-end distances. Note the very wide variety of possible configurations, the quite wide distribution of RMS end-to-end distances in the histogram, and the large number of 'data' you have to accumulate to generate a representative distribution. Note also that there is a most probable RMS end-to-end distance and that this is the square root of the fully extended length of the polymer, i.e. 10001/2. Since this end-to-end distance is the most probable, it is the one for which the entropy has its highest value.
(ii) The lever changes the polymer as follows. It incorporates a number of the initial segments into a single segment (changes the persistence length) and correspondingly reduces the number of segments, thus increasing the length of an individual segment while keeping the fully extended length of the polymer constant. Set the value of the slider to a segment length of 2. Press restart to restrt the calculation of the histogram. You will notice that the values of the RMS end-to-end distance increase (the scales in the left hand graph have been kept as the original segment length of unity). Repeat the calculation until you have a representative histogram and compare the RMS end-to-end distances for segment lengths 1 and 2. The ratio should be 1.4 (21/2). The applet does the calculation numerically but the RMS end-to-end distance can also be related to the molecular weight and the length of the individual segments analytically as described below. The random flight model only seems unrealistic if one makes the mistake of assuming that a random flight segment is the same as a chemical segment. For each homopolymer the number of chemical segments needed to produce a segment that follows the random flight model varies according to the stiffness of the structure and some values are given for common polymers below. This is essentially the calculation (ii) that you made for the applet above. The freely jointed chain consists of segments (not necessarily the chemical
segments) attached in such a way that each segment may take up any
orientation, with equal probability, with respect to its nearest neighbour,
i.e. if lk is a vector representing segment k then
and
for all values of j ≠ k and where τ is the angle between segments
k and j. A possible measure of the dimensions of the polymer is the end to end distance,
R. The problem is exactly the same as a random walk. After a
large number of steps the walker on average ends up exactly where he
started, i.e. <R> = 0. This is because
negative (backward) steps are as likely as positive (forward) ones.
However, the mean square distance,<R2>, is not zero and
that is what we take to characterize the chain length.
Thus
where N is the total number of segments, M is the molecular weight, and
m is the segment molecular weight. The important result is that the root
mean square end to end distance is proportional to the square root of the
molecular weight as shown in the applet above.
The random flight molecule is obviously unrealistic in that adjacent chemical segments cannot rotate freely. However, it is reasonable to divide the chain into sequences of chemical segments that are sufficiently long that these sequences obey the random flight molecule. If there are s chemical segments of molecular weight m in such a sequence the following result is obtained
The value of s is then a measure of the rigidity of the polymer. Some values are given below.
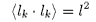
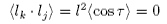
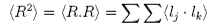
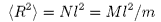
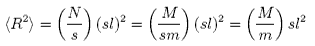
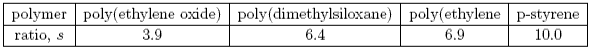
Dimensions of a polymer in solution: experiment
Scattering of radiation (light, neutrons, x-rays) id used to measure polymer dimensions. These methods give the radius of gyration Rg. Rg is the root mean square separation between all pairs of segments in a polymer molecule. For a random flight polymer 6<Rg2> = <Ro2> where Ro is the root mean square end-to-end distance.
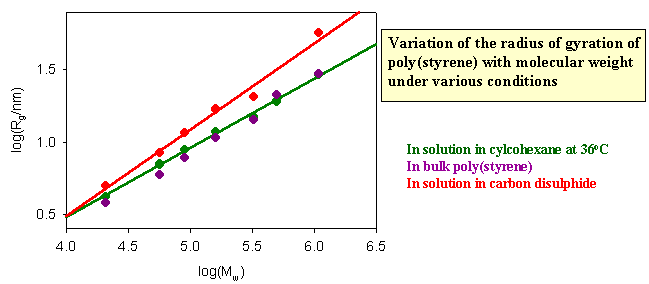
The dimensions of a polymer in solution depend on the nature of the solution. If the nett interaction between distant chain segments is zero (this is the θ condition) the conformation should be that of the random flight model. The graph below shows results for different molecular weight poly(styrene)s in cyclohexane, for which the θ temperature is 360C. If the radius of gyration is correctly predicted by the random flight model random the slope of the plot of logRg against logMw should be 1/2, which it as at 360C. There is no nett interaction between segments in a θ solvent, where the free energy of penetration of the polymer by solvent is exactly balanced by the free energy of stretching. In a good solvent, however, there is a nett repulsion between segments because the solvent causes the polymer to expand because of the additional favourable enthalpy of interaction between segments and solvent. The radius of gyration then increases. Theoretically and experimentally it is found that <Rg2> is proportional to M6/5 in a good solvent. Carbon disulphide is a good solvent for poly(styrene) and the corresponding plot verifying the change in the behaviour is shown as the red line in the figure below. Also included in the figure are data for the dimensions of poly(styrene) in the bulk amorphous polymer. They are consistent with the random flight model.
The methods of using x-ray or neutron scattering or, if the polymer is sufficiently large, light scattering are described separately in Scattering Methods.
A completely different type of method for estimating polymer size is Gel permeation chromatography. This uses a chromatographic column to separate particles according to their effective size in solution. The solution is passed through a column of a rigid gel containing pores of a chosen size. Small molecules diffuse in and out of the pores and hence pursue a tortuous route and take a long time to elute through the column. The bigger the polymer molecule in the solution the less easily it will be trapped in one of the pores and the more quickly it will pass through the column. The refractive index (or any suitable physical property) of the solution coming out of the end of the column is monitored and the presence of polymer is easily detected. The relation between elution time and molecular weight is not linear and depends on a number of factors, one of which is the relation between mean radius and molecular structure of the polymer.
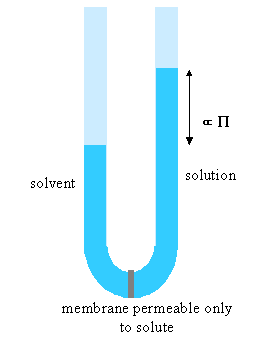
Osmotic pressure is the most useful of the colligative properties. It is widely used for the determination of molecular weights of large molecules such as polymers and proteins. The principle is shown in the diagram below. The pure solvent and solution are separated by a membrane permeable only to the small solvent molecules (the polymer cannot pass through). The chemical potential of the solvent is lower in the solution and it will therefore tend to pass through the membrane from pure solvent to solution. This process can be opposed by applying a pressure (additional to atmospheric and called the osmotic pressure) to the solution.
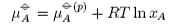
The effect of pressure on the chemical potential is determined as follows. From the first and second laws
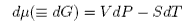
and, taking the temperature to be constant,
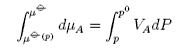
Hence
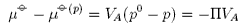
where Π is defined as the osmotic pressure. Hence,
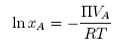
or, using
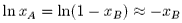
we obtain
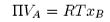
Note that the units of VA must be in m3 mol−1. If the concentration of polymer, cB is in kg m−3 ( = g dm−3),
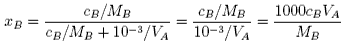
Hence
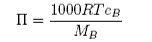
The osmotic effect is large and measurement of osmotic pressure is widely used for determining the molecular weights of polymers and proteins. For a typical large molecule as solute which produces a vapour pressure lowering of 10−2mmHg, the equivalent depression of freezing point will be 0.02 K, and the osmotic pressure will be 76 mmHg.
The thermodynamic properties of a polymer in solution are important in the determination of the molecular weight by osmotic pressure. Because a polymer is very large in comparison with the solvent molecule the entropy of mixing does not take on the simple form that it has in an ideal solution but has to be expressed in terms of volume fractions, i.e..
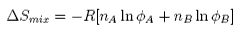
where φ are volume fractions. This leads to an entropic contribution to the chemical potential, which is
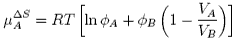
There is also a contribution to the mixing from the enthalpy of mixing of polymer and solvent. Any small enthalpic interaction between a polymer segment and a solvent has to be multiplied by the number of segments. Thus, even small interactions become significant. The calculation of the enthalpy of mixing is similar to that used in the regular solution model except that volume fractions are used instead of mole fractions, and the contribution is
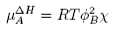
where χ is the mixing parameter (cf. β in the regular solution model). When these two contributions are used instead of the ideal contribution from mixing the result is
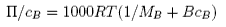
where B is
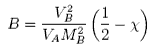
Comparison of the ideal and non-ideal expressions for the chemical potential shows that B contains one contribution from the excess entropy of mixing (the 1/2 term in the equation above) and one from the excess enthalpy of mixing (the c term). Taking the enthalpy of mixing to be zero leads to a positive value of B, resulting from the positive excess entropy of mixing. The positive excess entropy of mixing creates an additional driving force for diluting the solution with solvent, i.e. a higher osmotic pressure than would be expected from ideal behaviour. The enthalpy of mixing is proportional to the c parameter and is generally positive. A positive c makes mixing less favourable and hence lowers the osmotic pressure below the ideal value. The osmotic pressure appropriate to ideal mixing is obtained by putting B = 0. B is therefore analogous to the second virial coefficent for a gas, for which B is zero when the repulsion arising from the finite size of the molecules is exactly compensated by the van der Waals attraction. For a gas this happens at the Boyle temperature, at which the gas behaves as though it were ideal. This suggests a similar interpretation for the polymer solution. When B is positive the segments behave as though they repel one another because the solvent is forcing them apart (good solvent). When B is negative the positive enthalpy of mixing (c must now be greater than 1/2) makes the solvent a poorer solvent and hence the segments effectively attract one another. At the q point, B is zero and there is no nett interaction between polymer segments. It is at this point that the dimensions of the polymer should conform to the random flight model. Since the entropic contribution to the free energy varies with temperature, the q point can only be at one temperature. However, the quality of a mixed solvent can also be adjusted by varying its composition to give a q point dependent on composition (and temperature).
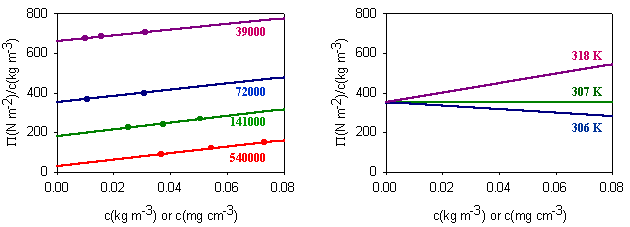
Measurement of osmotic pressure is useful for determining the molecular weight of a polymer. The equation indicates that a plot of P/c against c will give a straight line graph with intercept 1000RT/M2 and a slope proportional to B. Since B is positive for a good solvent the slope of the plot will be positive. Some actual data for different molecular weight samples of neoprene in toluene are shown on the left hand side of the figure below. The virial coefficient is the same for all molecular weights so the slopes are all the same. On the right hand side is shown the effect of varying temperature for a sample of poly(styrene) in cyclohexane, for which the q point is 340C. B is positive at higher temperatures and negative at lower temperatures. Since cyclohexane is a poor solvent for poly(styrene) below 340C one would expect the solubility to be reduced in some way. In fact, the polymer will become insoluble either by a further lowering of the temperature or by an increase in molecular weight.
The molecules in a synthetic polymer sample have a spread of molecular weights. Any measurement of molecular weight therefore gives an average. Different physical techniques measure different averages. Osmotic pressure, being a colligative property, gives the number average molecular weight, i.e.
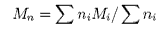
where ni is the number of molecules of weight Mi. An alternative method of measuring the molecular weight is to use light scattering (see below), which gives the weight average molecular weight, defined by
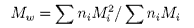
It is important point to note that, because polymer samples consist of mixtures of different molecular weight molecules, Mn and Mw are not the same. The extent to which they differ depends on the polydispersity of the sample (the ratio is often used as a measure of the polydispersity; ideally it should be as close to unity as possible). The procedure for determining the molecular weight of a polymer is to calibrate the column using known, monodisperse samples with structures as similar as possible to the sample of interest. Both Mn and Mw can be evaluated from the shape of the elution peak.
There are many good books on Polymers but they mostly cover far more ground than is needed for the course and so only a small number are listed here. The Colloidal Domain by D.F.Evans and H.Wennerstrom (VCH) covers all aspects of the course. The most appropriate quantitative descriptions are in Polymer Physics by U.W.Gedde, Chapman & Hall. A good, basically qualitative, book on polymers is Giant Molecules by A.Y.Grosberg and A.R.Khokhlov (Academic Press). Other useful books on polymers at this level are Sperling: Introduction to Physical Polymer Science, Billmeyer: Textbook of Polymer Science, Cowie: Introduction to Polymer Science. A good discussion of the thermodynamics of solutions, including polymer solutions, is Ch. 10 of Caldin: Chemical Thermodynamics, but it is out of print and not easy to obtain.